The goal of squat is to provide extensions of common statistical methods for the analysis of unit quaternion time series. Available statistical methods for QTS samples are currently:
rnorm_qts(),scale(),mean(),median(),dist()
including the possibility of separating amplitude and phase
variability,prcomp(),kmeans().You can install the official version from CRAN via:
install.packages("squat")or you can opt to install the development version from GitHub with:
# install.packages("devtools")
devtools::install_github("LMJL-Alea/squat")library(squat)First, let us visualize the sample of QTS from the
vespa64 dataset included in the package. The package
provides two ways of doing this: either via a static plot or via an
animated one (which uses gganimate behind the
scenes and will prompt you to install it in case you have not
already).
Here is the static version:
plot(vespa64$igp)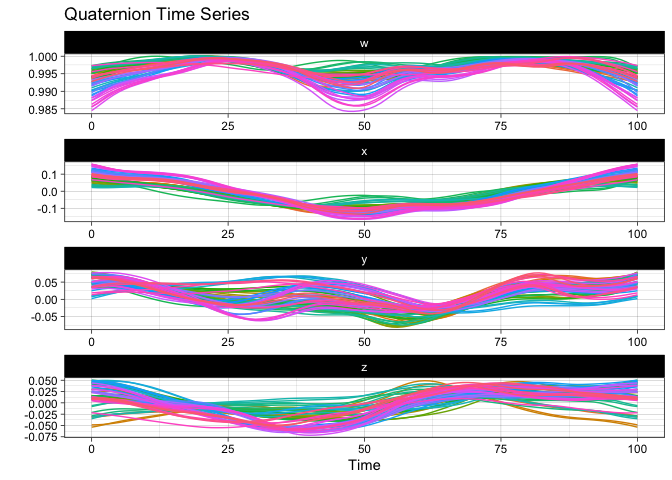
You can also use ggplot2::autoplot() instead of
plot() to save the resulting ggplot object for
further customization.
Here is the animated version:
p <- ggplot2::autoplot(vespa64$igp, with_animation = TRUE)
gganimate::anim_save("man/figures/README-animated-plot.gif", p)You can compute the geometric mean of the sample and append it to the sample for visualization:
m <- mean(vespa64$igp)
sample_and_mean <- append(vespa64$igp, m)
plot(sample_and_mean, highlighted = c(rep(FALSE, 64), TRUE))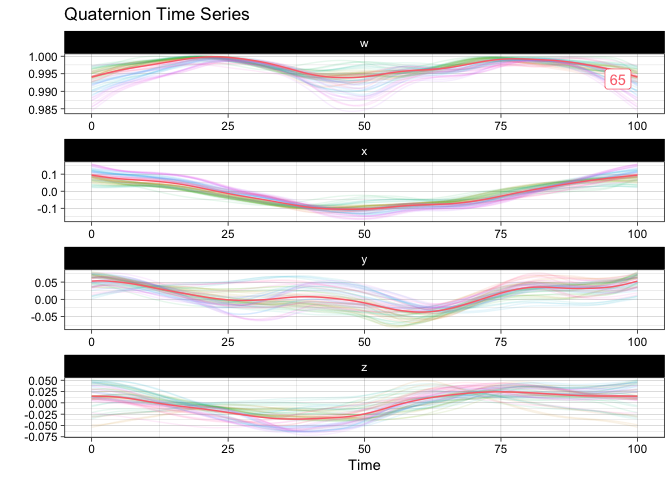
You can compute the pairwise distance matrix (based on the DTW for now):
D <- dist(
vespa64$igp,
is_domain_interval = TRUE,
transformation = "srvf",
metric = "l2",
warping_class = "bpd"
)
C <- exp(-D / (sqrt(2) * sd(D)))
C <- (C - min(C)) / diff(range(C))
C <- C |>
as.matrix() |>
corrr::as_cordf()
corrr::network_plot(C)
#> Warning: ggrepel: 1 unlabeled data points (too many overlaps). Consider
#> increasing max.overlaps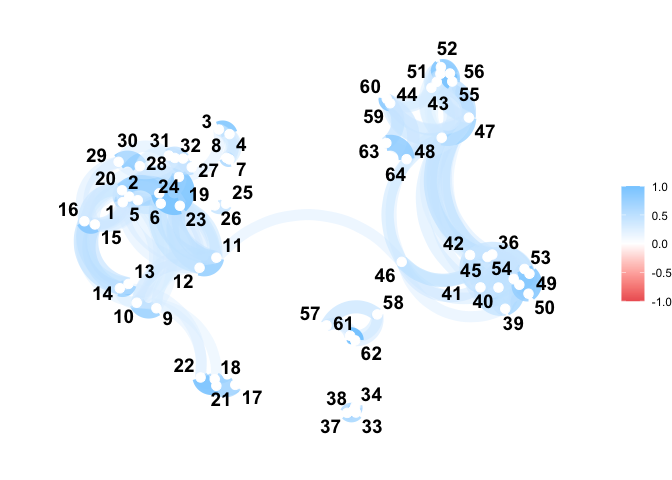
You can perform tangent principal component analysis and visualize it:
tpca <- prcomp(vespa64$igp)
#> ℹ The maximum number of principal component is 63.
plot(tpca, what = "PC1")
#> The `original_space` boolean argument is not specified. Defaulting to TRUE.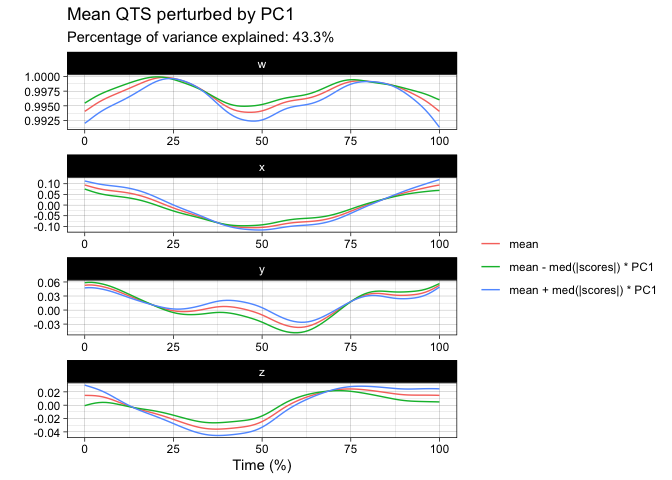
plot(tpca, what = "scores")
#> The `plane` length-2 integer vector argument is not specified. Defaulting to
#> 1:2.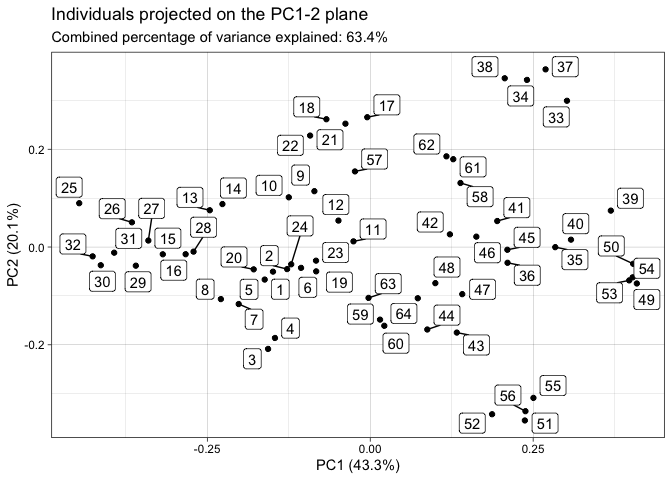
screeplot(tpca)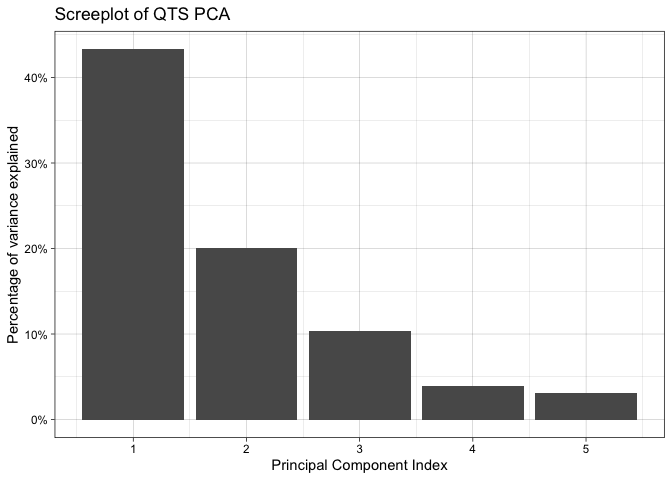
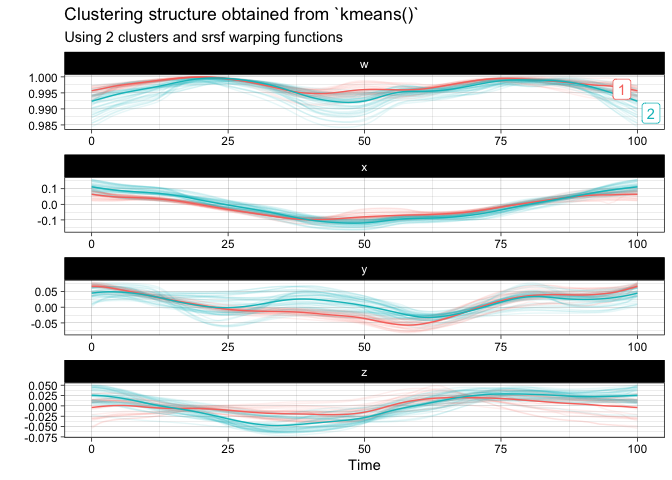
You can finally perform a k-means clustering and visualize it:
km <- kmeans(
vespa64$igp,
n_clusters = 2,
is_domain_interval = TRUE,
transformation = "srvf",
warping_class = "bpd"
)
plot(km)